RADHIKA NAIR, PhD
PhD National Institute of Immunology, New Delhi
Career Development Fellow, University of Cambridge, UK
Senior Research Officer, Garvan Institute of Medical Research, Australia
Conjoint lecturer, Faculty of Medicine, St Vincent’s Clinical School
Ramanujan Faculty Fellow, Rajiv Gandhi Centre for Biotechnology
Faculty, Centre for Human Genetic
Brief research summary of the laboratory:
Metastasis or the spread of cancer from the primary site to other parts of the body is a silent killer in breast cancer, with 80% mortality rate for women with metastatic disease. Key questions that need to be dealt with include heterogeneity and a detailed understanding of the cells critical for actual metastatic establishment. Therapeutic targeting of metastasis requires a deeper understanding of the metastatic process and our work will directly address key challenges in the field.
Comprehending the cell intrinsic mechanisms that allow a tumour cell to survive, remain in a state of dormancy and then thrive in a hostile new environment of a distant metastatic organ is vital. In addition, analyzing the role of the extracellular environment in supporting metastatic cells is key to giving us a complete picture of the metastatic process. The hypothesis underpinning the work in my laboratory is that breast cancer cells form metastases utilizing a combination of cell autonomous (‘intrinsic’) programs and microenvironmental (‘extrinsic’) changes. This work has implications for conceptually understanding the critical molecules involved in the metastatic process and identifying its “Achilles’ heel”, which can be exploited for therapeutic purposes.
Approach 1- Investigating the role of known molecular mediators of metastasis in breast cancer
I have focused on the Inhibitor of differentiation (or Id) family of bHLH transcriptional repressor proteins which play a critical role in the metastatic spread of breast cancer cells to distant organs especially the lung. I have gone onto identify two potential pathways by which Id proteins control key cancer phenotypes – via negative transcriptional regulation of the Robo1 pathway and cell cycle pathway by impacting Kif11 and AurkA ( see Publications).
Approach 2: Identifying the molecular mediators involved in breast cancer metastasis using an unbiased approach
I plan to expand this work to identify pathways which are critical for the metastatic phenotypes hardwired into the metastatic cells with the aim of discovering new potential therapeutic targets. I have isolated two phenotypically distinct tumor cell populations with differing metastatic potential. Resolving the molecular drivers behind the intratumoral heterogeneity revealed critical players which are druggable ( see Publications). Using a validation cohort of matched primary and metastatic tumors will allow me to further understand the genetics underlying the metastatic process and drive my work into more translational avenues in an Indian context.
Selected Publications:
- Sarah Hyder Iqbal, Krishnaveni Mishra, Radhika Nair, Rashna Bhandari, Sreelaja Nair and Suhita Nadkarni.
From Numbers to Narratives: Towards Gender Equity in Academia.
Dialogue - Science, Scientists, and Society (2026).
- Borker NS, Sajimon J, Subhadarshini S, Aara R, Jolly MK, Prabhu JS, Nair R. Role of intratumoral heterogeneity in metastatic progression and drug resistance.
Discover Oncology 2025;16:1689.
doi: 10.1007/s12672-025-03322-4
- Neilah Firzan CA*, Rifat Aara*, Kalyanashis Jana*, Sreelakshmi Radhakrishnan*, Mubeena S, Radhika Nair, Harsha Bajaj, Ulrich Kleinekathöfer, and Kozhinjampara R Mahendran. ( * co first authors, CHG)
Fabrication of cytotoxic mirror image nanopores
Nature Communications 16, 8666 (2025).
doi: 10.1038/s41467-025-64025-6
- Repurposing Statins in Combination Therapy for effective ablation of metastatic breast cancer
Rifat Aara, Neeha Sinai Borker, Jyothilakshmi Sajimon, Athul Krishnan R, Mohit Kumar Jolly, Jyothi S Prabhu, Radhika Nair.
Advances in Cancer Biology - Metastasis,
Volume 16, 2026 doi: 10.1016/j.adcanc.2025.100166
- Phenotypic heterogeneity drives differential disease outcome in Triple Negative Breast Cancer
Archana P. Thankamony, Reshma Murali, Priyanka Chakraborty, Nitheesh Karthikeyan, Binitha Anu Varghese, Vishnu Sunil Jaikumar, Mohit Kumar Jolly and Radhika Nair
Front. Oncol. Sec. Cancer Molecular Targets and Therapeutics
Volume 13 - 2023 | doi: 10.3389/fonc.2023.1230647
- Unravelling the dangerous Duet between Cancer cell plasticity and drug resistance.
Chatterjee, Namrata; Pulipaka, Bhavana ; Subbalakshmi, Ayalur Raghu ; Jolly, Mohit Kumar; Nair , Radhika
July 2023, Computational and Systems Oncology; https://doi.org/10.1002/cso2.1051
- Lineage Plasticity in Cancer: The Tale of a Skin-walker.
Thankamony, A.P.; Subbalakshmi, A.R.; Jolly, M.K.; Nair, R.
Cancers 2021, 13(14), 3602; https://doi.org/10.3390/cancers13143602
- Archana PT$, Kritika Saxena$ ,Reshma Murali, Mohit K Jolly*, Radhika Nair*
$ Co corresponding authors
Cancer Stem Cell plasticity: a deadly deal
(Invited review, Frontiers in Molecular Biosciences)
Front. Mol. Biosci., 30 April 2020 | https://doi.org/10.3389/fmolb.2020.00079
- Thankamony, A.P.; Murali, R.; Karthikeyan, N.; Varghese, B.A.; Teo, W.S.; McFarland, A.; Roden, D.L.; Holliday, H.; Konrad, C.V.; Cazet, A.; Dodson, E.; Yang, J.; Baker, L.A.; George, J.T.; Levine, H.; Jolly, M.K.; Swarbrick, A.; Nair, R* (*corresponding author).
Targeting the Id1-Kif11 Axis in Triple-Negative Breast Cancer Using Combination Therapy.
Biomolecules, 8 Sep 2020, 10, 1295.
- Wee S Teo, Aurélie S Cazet, Daniel L Roden, Nitheesh K, Holly Holliday,Christina Konrad, Reshma Murali , Binitha Anu Varghese, Archana PT, Eoin Dodson, Kate Harvey, Andrea McFarland, Simon Junankar, Sunny Ye, Jessica Yang, Iva Nikolic, Jaynish Shah, Laura A Baker, Ewan K.A.Millar, Mathew J Naylor, Christopher J Ormandy, Sunil R Lakhani, Warren Kaplan,Albert Mellick, Sandra A O’Toole, Alexander Swarbrick*, Radhika Nair*
(*Co corresponding and Co senior authorship)
Id proteins promote a cancer stem cell phenotype in mouse models of triple negative breast cancer via Robo1-dependent c-Myc activation
Front. Cell Dev. Biol., 17 July 2020 | https://doi.org/10.3389/fcell.2020.00552
- Laura A Baker, Christoph Krisp, Daniel Roden, Holly Holliday, Sunny Z Wu, Simon Junankar, Aurelien A Serandour, Hisham Mohammed, Radhika Nair, ChiaLing Chan, Jessica Yang, Nicola Foreman, Breanna Fitzpatrick, Geetha Sankaranarayanan, Andrew MK Law, Chris Ormandy, MatthewJ Naylor, Andrea McFarland, Peter T Simpson, Sunil Lakhani, Sandra O'Toole, Christina Selinger, Lyndal Anderson, Goli Samimi, Neville F Hacker, Warren Kaplan, Jason S Carroll, Mark Molloy, Alexander Swarbrick.
Proteogenomic analysis of Inhibitor of Differentiation 4 (ID4) in basal-like breast cancer.
Breast Cancer Res 22,63 (2020). https://doi.org/10.1186/s13058-020-01306-6
- Aurélie Cazet*, Mun Hui*, Ben Elsworth, Sunny Wu, Caroline Cooper, Michael Samuel, Jessica Yang, Niantao Deng, Nicola Foreman, Andrea McFarland, Radhika Nair, Sandra O’Toole, Rosalía Caballero, Miguel Martín, Alexander Swarbrick.
Stromal Smoothened inhibition depletes triple negative breast cancer stem cells and sensitizes to chemotherapy.
Nature Communications. 2018 Jul 24;9(1):2897. doi: 10.1038/s41467-018-05220-6.
- Konrad C, Murali R, Varghese AB, Nair R* (* corresponding author).
The role of cancer stem cells in tumor heterogeneity and resistance to therapy.
Can J Physiol Pharmacol. 2017 Jan;95(1):1-15. doi: 10.1139/cjpp-2016-0079.
- Teo SW, Nair R, Swarbrick A.
New insights into the role of ID proteins in breast cancer metastasis: a MET affair.
Breast Cancer Res. 2014 May 15;16(2):305. doi: 10.1186/bcr3654.
- Nair R*, Teo SW*, Mittal V, Swarbrick A.
ID proteins regulate diverse aspects of cancer progression and provide novel therapeutic oppurtunities (* co-first authors).
Mol Ther. 2014 Aug;22(8):1407-1415. doi: 10.1038/mt.2014.83. Epub 2014 May 14.
- PimentelB*-Nair R*, Bermejo-Rodriguez C, Agu C, Preston M, Wang X,Bernal JA, Sherrat D, de la Cueva-Mendez G. (*co-first authors).
Toxin Kid uncouples DNA replication and cell division to enforce retention of plasmid R1 in E.coli cells .
Proc Natl Acad Sci U S A. 2014 Feb 18;111(7):2734-9. doi: 10.1073/pnas.1308241111.
- Nair R, Roden DL, Teo WS, McFarland A, Junankar S , Ye S,Nguyen A, Yang J, Nikolic I, Morey A, Shah J, Selinger C, Baker LA, Cowley M, Naylor MJ, Ormandy CJ, Lakhani SR, Kaplan W, O’Toole SA, Swarbrick, A.
c-Myc and Her2 cooperate to drive a stem-like phenotype with poor prognosis in breast cancer
Oncogene. 2014;33(30):3992-4002. doi:10.1038/onc.2013.368
- Invited to be the Topic Editor for Special Issue on “Multifaceted dynamics of Metastasis” in Frontiers in Cell and Developmental Biology 2021
https://www.frontiersin.org/research-topics/20087/multifaceted-dynamics-of-metastasis#overview
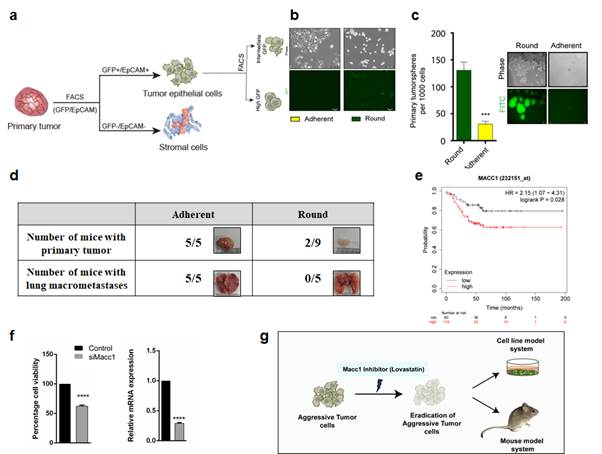
Isolation of phenotypically distinct tumor cells from a heterogeneous population.
(a,b) Schematic showing the sorting of primary tumor. (c) Tumorsphere assay. (d) The 4T1Adherent cells are highly aggressive. (e) Relevance of Macc1 to breast cancer (Kaplan-Meier survival curve).(f) Knock down of Macc1 lead to significant loss in proliferation of T1 aggressive cells. (g) Schematic for therapeutic targeting of Macc1 using small molecule inhibitors.
Data are expressed as mean ± standard deviation. n=3, p <0.05 are considered statistically significant with *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001




Stay Up to Date With Whats Happening